This article is a continuation of previous articles on deep neural network and predictor selection. Here we will cover features of a neural network initiated by Stacked RBM, and its implementation in the “darch” package. The possibility of using a hidden Markov model for improving the performance of a neural network prediction will also be revealed. In conclusion, we will programmatically implement an operational Expert Advisor.
Contents
- 1. Structure of DBN
- 2. Preparation and selection of data
- 2.1. Input variables
- 2.2. Output variables
- 2.3. Initial data frame
- 2.3.1. Deleting highly correlated variables
- 2.4. Selection of the most important variables
- 3. Experimental part.
- 3.1. Building models
- 3.1.1. Brief description of the “darch” package
- 3.1.2. Building the DBN model. Parameters.
- 3.2. Formation of training and testing samples.
- 3.2.1. Balancing classes and pre-processing.
- 3.2.2. Coding the target variable
- 3.3. Training the model
- 3.3.1. Pre-training
- 3.3.2. Fine-tuning
- 3.4. Testing the model. Мetrics.
- 3.4.1. Decoding predictions.
- 3.4.2. Improving the prediction results
- Calibration
- Smoothing with a Markov chain model
- Correcting predicted signals on the theoretical balance curve
- 3.4.3. Metrics
- 3.1. Building models
- 4. Structure of the Expert Advisor
- 4.1. Description of the Expert Advisor’s operation
- 4.2. Self-control. Self-training
- Installation and launching
- Ways and methods of improving qualitative indicators.
- Conclusion
Introduction
In preparation of data for conducting experiments, we will use variables from the previous article about evaluating and selecting predictors. We will form the initial sample, clean it and select the important variables.
We will consider ways of dividing the initial sample into training, testing and validation samples.
Using the “darch” package we will build a model of the DBN network, and train it on our sets of data. After testing the model, we will obtain metrics that will enable us to evaluate quality of the model. We will consider many opportunities that the package offers to configure settings of a neural network.
Also, we will see how hidden Markov models can help us improve neural network predictions.
We will develop an Expert Advisor where a model will be trained periodically on the fly without interruption in trade, based on results of continuous monitoring. The DBN model from the “darch” package will be used in the Expert Advisor. We will also incorporate the Expert Advisor built using SAE DBN from the previous article.
Furthermore, we will indicate ways and methods of improving qualitative indicators of the model.
1. Structure of a deep neural network initialized by Stacked RBM (DN_SRBM)
I recall that DN_SRBM consists of n-number of RBM that equals the number of hidden layers of neural network and, basically, the neural network itself. Training comprises two stages.
The first stage involves PRE-TRAINING. Every RBM is systematically trained without a supervisor on the input set (without target). After this weight of hidden layers, RBM are transferred to relevant hidden layers of neural network.
The second stage involves FINE-TUNING, where neural network is trained with a supervisor. Detailed information about it was provided in the previous article, so we don’t have to repeat ourselves here. I will simply mention that unlike the “deepnet” package that we have used in the previous article, the “darch” package helps us to implement wider opportunities in building and tuning the model. More details will be provided when creating the model. Fig. 1 shows the structure and the training process of DN_SRBM
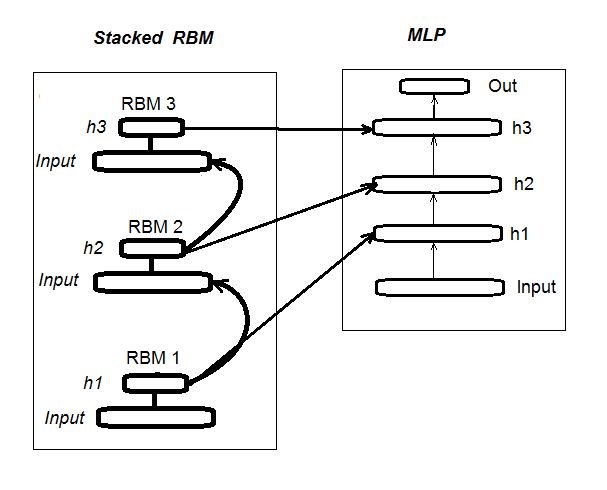
Fig. 1. Structure of DN SRBM
2. Preparation and selection of data
2.1. Input variables (signs, predictors)
In the previous article, we have already considered the evaluation and selection of predictors, so there is no need to provide additional information now. I will only mention that we used 11 indicators (all oscillators: ADX, aroon, ATR, CCI, chaikinVolatility, CMO, MACD, RSI, stoch, SMI, volatility). Several variables were selected from some indicators. This way we have formed the input set of 17 variables. Let’s take quotes from the last 6000 bars on EURUSD, М30 as at 14.02.16, and calculate indicator values using the In() function.
#---2--------------------------------------------- In <- function(p = 16){ require(TTR) require(dplyr) require(magrittr) adx <- ADX(price, n = p) %>% as.data.frame %>% mutate(.,oscDX = DIp - DIn) %>% transmute(.,DX, ADX, oscDX) %>% as.matrix() ar <- aroon(price[ ,c('High', 'Low')], n = p) %>% extract(,3) atr <- ATR(price, n = p, maType = "EMA") %>% extract(,1:2) cci <- CCI(price[ ,2:4], n = p) chv <- chaikinVolatility(price[ ,2:4], n = p) cmo <- CMO(price[ ,'Med'], n = p) macd <- MACD(price[ ,'Med'], 12, 26, 9) %>% as.data.frame() %>% mutate(., vsig = signal %>% diff %>% c(NA,.) %>% multiply_by(10)) %>% transmute(., sign = signal, vsig) %>% as.matrix() rsi <- RSI(price[ ,'Med'], n = p) stoh <- stoch(price[ ,2:4], nFastK = p, nFastD =3, nSlowD = 3, maType = "EMA") %>% as.data.frame() %>% mutate(., oscK = fastK - fastD) %>% transmute(.,slowD, oscK) %>% as.matrix() smi <- SMI(price[ ,2:4],n = p, nFast = 2, nSlow = 25, nSig = 9) kst <- KST(price[ ,4])%>% as.data.frame() %>% mutate(., oscKST = kst - signal) %>% select(.,oscKST) %>% as.matrix() In <- cbind(adx, ar, atr, cci, chv, cmo, macd, rsi, stoh, smi, kst) return(In) }
We will get the input data matrix on the output.
2.2 Output data (target variable)
As a target variable we take signals obtained with ZZ. The function calculating a zigzag and a signal:
#----3------------------------------------------------ ZZ <- function(pr = price, ch = ch , mode="m") { require(TTR) require(magrittr) if (ch > 1) ch <- ch/(10 ^ (Dig - 1)) if (mode == "m") {pr <- pr[ ,'Med']} if (mode == "hl") {pr <- pr[ ,c("High", "Low")]} if (mode == "cl") {pr <- pr[ ,c("Close")]} zz <- ZigZag(pr, change = ch, percent = F, retrace = F, lastExtreme = T) n <- 1:length(zz) dz <- zz %>% diff %>% c(., NA) sig <- sign(dz) for (i in n) { if (is.na(zz[i])) zz[i] = zz[i - 1]} return(cbind(zz, sig)) }
Function parameters:
pr = price – matrix of OHLCMed quotes;
ch – minimum length of the zigzag bend in the points (4 signs) or in real terms (for example, ch = 0.0035);
mode – applied price (“m” – medium, “hl” – High and Low, “cl” – Close), medium used by default.
The function returns the matrix with two variables — in fact, the zigzag and the signal, obtained on the base of the zigzag angle in the range of [-1;1]. We shift the signal by one bar to the left (towards future). This specific signal will be used to train the neural network.
We calculate signals for ZZ with a bend length of at least 37 points (4 signs).
> out <- ZZ(ch = 37, mode = "m") Loading required package: TTR Loading required package: magrittr > table(out[ ,2]) -1 1 2828 3162
As we can see, the classes are slightly unbalanced. When forming samples for training the model, we will take necessary measures to level them off.
2.3. Initial data frame
Let’s write a function that will create the initial data frame, clean it from uncertain data (NA) and convert the target variable to the factor with two classes “-1” and “+1”. This function combines previously written functions In() and ZZ(). We will instantly crop the last 500 bars that will be used to evaluate the quality of the model’s prediction.
#-----4--------------------------------- form.data <- function(n = 16, z = 37, len = 500){ require(magrittr) x <- In(p = n) out <- ZZ(ch = z, mode = "m") data <- cbind(x, y = out[ ,2]) %>% as.data.frame %>% head(., (nrow(x)-len))%>% na.omit data$y <- as.factor(data$y) return(data) }
2.3.1. Deleting highly correlated variables
We will delete variables with a correlation coefficient above 0.9 from our initial set. We will write a function that will form the initial data frame, remove highly correlated variables and return clean data.
We can check in advance which variables have a correlation above 0.9.
> data <- form.data(n = 16, z = 37) # prepare data frame > descCor <- cor(data[ ,-ncol(data)])# remove a target variable > summary(descCor[upper.tri(descCor)]) Min. 1st Qu. Median Mean 3rd Qu. Max. -0.1887 0.0532 0.2077 0.3040 0.5716 0.9588 > highCor <- caret::findCorrelation(descCor, cutoff = 0.9) > highCor [1] 12 9 15 > colnames(data[ ,highCor]) [1] "rsi" "cmo" "SMI"
Thus, the above listed variables are subject to removal. We will delete them from the data frame.
> data.f <- data[ ,-highCor] > colnames(data.f) [1] "DX" "ADX" "oscDX" "ar" "tr" [6] "atr" "cci" "chv" "sign" "vsig" [11] "slowD" "oscK" "signal" "vol" "Class"
We will write it compactly in one function:
#---5----------------------------------------------- cleaning <- function(n = 16, z = 37, cut = 0.9){ data <- form.data(n, z) descCor <- cor(data[ ,-ncol(data)]) highCor <- caret::findCorrelation(descCor, cutoff = cut) data.f <- data[ ,-highCor] return(data.f) } > data.f <- cleaning()
Not all authors of packages and researchers agree that highly correlated data should be removed from the sets. However, results using both options should be compared here. In our case, we will select the option with deleting.
2.4. Selection of the most important variables
Important variables will be selected based on three indicators: global importance, local importance (in conjunction) and partial importance by class. We will seize the opportunities of the “randomUniformForest” package as detailed in the previous article. All previous and following actions will be gathered in one function for compactness. Once executed, we will obtain three sets as a result:
- with best variables in contribution and interaction;
- with best variables for the class “-1”;
- with best variables for the class “+1”.
#-----6------------------------------------------------ prepareBest <- function(n, z, cut, method){ require(randomUniformForest) require(magrittr) data.f <<- cleaning(n = n, z = z, cut = cut) idx <- rminer::holdout(y = data.f$Class) prep <- caret::preProcess(x = data.f[idx$tr, -ncol(data.f)], method = method) x.train <- predict(prep, data.f[idx$tr, -ncol(data.f)]) x.test <- predict(prep, data.f[idx$ts, -ncol(data.f)]) y.train <- data.f[idx$tr, ncol(data.f)] y.test <- data.f[idx$ts, ncol(data.f)] #--------- ruf <- randomUniformForest( X = x.train, Y = y.train, xtest = x.test, ytest = y.test, mtry = 1, ntree = 300, threads = 2, nodesize = 1 ) imp.ruf <- importance(ruf, Xtest = x.test) best <- imp.ruf$localVariableImportance$classVariableImportance %>% head(., 10) %>% rownames() #-----partImport best.sell <- partialImportance(X = x.test, imp.ruf, whichClass = "-1", nLocalFeatures = 7) %>% row.names() %>% as.numeric() %>% colnames(x.test)[.] best.buy <- partialImportance(X = x.test, imp.ruf, whichClass = "1", nLocalFeatures = 7) %>% row.names() %>% as.numeric() %>% colnames(x.test)[.] dt <- list(best = best, buy = best.buy, sell = best.sell) return(dt) }
We will clarify the order of the function calculations. Official parameters:
n – input data parameter;
z – output data parameter;
cut – correlation threshold of variables;
method – input data pre-processing method.
Order of calculations:
- create the initial set of data.f, which has highly correlated variables removed, and save it for further use;
- identify indexes of the training and testing samples of idx;
- determine pre-processing parameters of prep;
- divide the initial sample into training and testing samples, input data normalized;
- obtain and test the ruf model on the obtained sets;
- calculate the importance of the imp.ruf variables;
- select 10 most important variables in terms of contribution and interaction — best;
- select 7 most important variables for each class “-1” and “+1” — best.buy, best.sell;
- Create a list with three sets of predictors — best, best.buy, best.sell.
We will calculate these samples and evaluate values of global, local and partial importance of the selected variables.
> dt <- prepareBest(16, 37, 0.9, c("center", "scale","spatialSign")) Loading required package: randomUniformForest Labels -1 1 have been converted to 1 2 for ease of computation and will be used internally as a replacement. 1 - Global Variable Importance (14 most important based on information gain) : Note: most predictive features are ordered by 'score' and plotted. Most discriminant ones should also be taken into account by looking 'class' and 'class.frequency'. variables score class class.frequency percent 1 cci 4406 -1 0.51 100.00 2 signal 4344 -1 0.51 98.59 3 ADX 4337 -1 0.51 98.43 4 sign 4327 -1 0.51 98.21 5 slowD 4326 -1 0.51 98.18 6 chv 4296 -1 0.52 97.51 7 oscK 4294 -1 0.52 97.46 8 vol 4282 -1 0.51 97.19 9 ar 4271 -1 0.52 96.95 10 atr 4237 -1 0.51 96.16 11 oscDX 4200 -1 0.52 95.34 12 DX 4174 -1 0.51 94.73 13 vsig 4170 -1 0.52 94.65 14 tr 4075 -1 0.50 92.49 percent.importance 1 7 2 7 3 7 4 7 5 7 6 7 7 7 8 7 9 7 10 7 11 7 12 7 13 7 14 7 2 - Local Variable importance Variables interactions (10 most important variables at first (columns) and second (rows) order) : For each variable (at each order), its interaction with others is computed. cci slowD atr tr DX atr 0.1804 0.1546 0.1523 0.1147 0.1127 cci 0.1779 0.1521 0.1498 0.1122 0.1102 slowD 0.1633 0.1375 0.1352 0.0976 0.0956 DX 0.1578 0.1319 0.1297 0.0921 0.0901 vsig 0.1467 0.1209 0.1186 0.0810 0.0790 oscDX 0.1452 0.1194 0.1171 0.0795 0.0775 tr 0.1427 0.1168 0.1146 0.0770 0.0750 oscK 0.1381 0.1123 0.1101 0.0725 0.0705 sign 0.1361 0.1103 0.1081 0.0704 0.0685 signal 0.1326 0.1068 0.1045 0.0669 0.0650 avg1rstOrder 0.1452 0.1194 0.1171 0.0795 0.0775 vsig oscDX oscK signal ar atr 0.1111 0.1040 0.1015 0.0951 0.0897 cci 0.1085 0.1015 0.0990 0.0925 0.0872 slowD 0.0940 0.0869 0.0844 0.0780 0.0726 DX 0.0884 0.0814 0.0789 0.0724 0.0671 vsig 0.0774 0.0703 0.0678 0.0614 0.0560 oscDX 0.0759 0.0688 0.0663 0.0599 0.0545 tr 0.0733 0.0663 0.0638 0.0573 0.0520 oscK 0.0688 0.0618 0.0593 0.0528 0.0475 sign 0.0668 0.0598 0.0573 0.0508 0.0455 signal 0.0633 0.0563 0.0537 0.0473 0.0419 avg1rstOrder 0.0759 0.0688 0.0663 0.0599 0.0545 chv vol sign ADX avg2ndOrder atr 0.0850 0.0850 0.0847 0.0802 0.1108 cci 0.0824 0.0824 0.0822 0.0777 0.1083 slowD 0.0679 0.0679 0.0676 0.0631 0.0937 DX 0.0623 0.0623 0.0620 0.0576 0.0881 vsig 0.0513 0.0513 0.0510 0.0465 0.0771 oscDX 0.0497 0.0497 0.0495 0.0450 0.0756 tr 0.0472 0.0472 0.0470 0.0425 0.0731 oscK 0.0427 0.0427 0.0424 0.0379 0.0685 sign 0.0407 0.0407 0.0404 0.0359 0.0665 signal 0.0372 0.0372 0.0369 0.0324 0.0630 avg1rstOrder 0.0497 0.0497 0.0495 0.0450 0.0000 Variable Importance based on interactions (10 most important) : cci atr slowD DX tr vsig oscDX 0.1384 0.1284 0.1182 0.0796 0.0735 0.0727 0.0677 oscK signal sign 0.0599 0.0509 0.0464 Variable importance over labels (10 most important variables conditionally to each label) : Class -1 Class 1 cci 0.17 0.23 slowD 0.20 0.09 atr 0.14 0.15 tr 0.04 0.12 oscK 0.08 0.03 vsig 0.06 0.08 oscDX 0.04 0.08 DX 0.07 0.08 signal 0.05 0.04 ar 0.04 0.02
Results
- In terms of global importance all 14 input variables are equal.
- The best 10 are defined by the overall contribution (global importance) and interaction (local importance).
- Seven best variables in partial importance for each class are shown on the charts below.
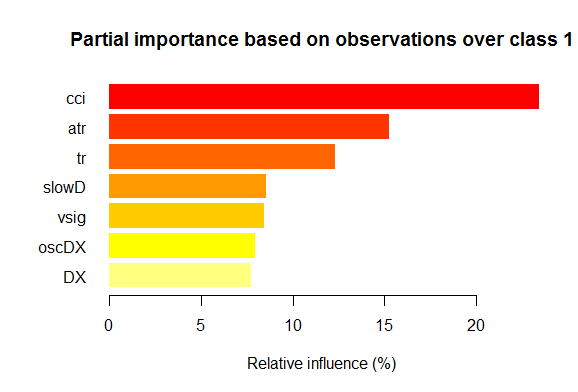
Fig. 2. Partial importance of variables for the “1” class
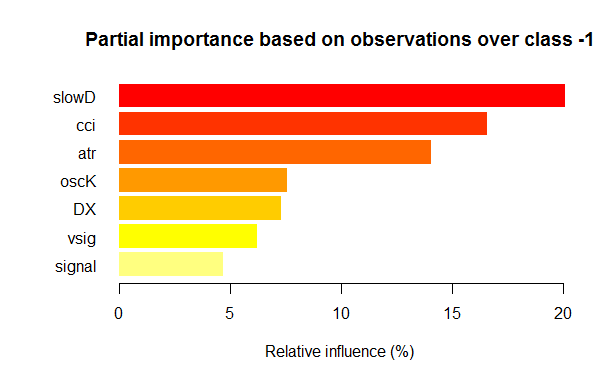
Fig. 3. Partial importance of variables for the “-1” class
As we can see, the most important variables for different classes are different in both structure and rankings. Thus, if for the “-1” class the slowD variable is the most important, then for the “+1” class it is only on the 4th place.
So, we have sets of data ready. Now we can proceed with the experiments.
3. Experimental part.
Experiments will be conducted in the R language — Revolution R Open, version 3.2.2, distribution of the Revolution Analytics company, to be specific. http://www.revolutionanalytics.com/revolution-r-open
This distribution has a number of advantages over regular R 3.2.2:
- quick and more qualitative calculations through applying the multi-threaded processing with Intel® Math Kernel Library ;
- advanced features of Reproducible R Toolkit. One slight clarification: the R language is actively developing by constantly improving the existing packages and adding the new ones. The flip side of such progress involves the loss of reproducibility. That is, your products that were written few months back and were functioning well, suddenly stop working after the next update of packages. Much time is wasted to identify and liquidate the error caused by the change in one of the packages. For example, the Expert Advisor attached to the first article on deep neural networks was functioning well at the point of creation. However, a few months after the publication a number of users have complained about its non-operability. The analysis showed that updating the “svSocket” package has led to the Expert Advisor’s malfunction, and I was unable to find the reason behind it. The finalized Expert Advisor will be attached to this article. This problem has become a pressing issue, and it was easily solved in Revolution Analytics. Now, when a new distribution is released, the condition of all packages in the CRAN repositary is fixed at the release date by copying them on their mirror. No changes in the CRAN depositary after this date can affect the packages “frozen” on the Revolution mirror. Furthermore, starting from October 2014, the company makes daily snapshots of the CRAN depositary, fixing the relevant state and versions of packages. With their own “checkpoint” package we can now download necessary packages that are relevant at the date we need. In other words, we operate a some kind of time machine.
And another news. Nine months ago, when Microsoft purchased Revolution Analytics, it promised to support their developments and kept the Revolution R Open (RRO) distribution available free of charge. It was followed by numerous messages about novelties in RRO and Revolution R Enterpise (not to mention the integration of R with SQL Server, PowerBI, Azure and Cortana Analitics). Now we have information that the next RRO update will be called Microsoft R Open, and Revolution R Enterprise — Microsoft R Server. And not so long ago Microsoft has announced that R will be available in Visual Studio. R Tools for Visual Studio (RTVS) follows the Python Tools for Visual Studio model. It will be a free addition to Visual Studio that will provide a complete IDE for R with the possibility to edit and debug the scripts interactively.
By the time the article was finished, Microsoft R Open (R 3.2.3) was already released, therefore, further in the article we will refer to packages for this version.
3.1. Building models
3.1.1. Brief description of the “darch” package
The “darch” ver. 0.10.0 package offers a wide range of functions that don’t just allow to create and train the model, but, literally, build it brick by brick and adjust it according to your preferences. As previously indicated, deep neural network consists of n-number of RBM (n = layers -1) and MLP neural network with a number of layers. Layer-wise pre-training of RBM is executed on unformatted data without a supervisor. Fine-tuning of neural network is performed with a supervisor on formatted data. Dividing the training stages gives us an opportunity to use data various in volume (but not structure!) or to obtain several various fine-tuned models on the basis of pre-training alone. Furthermore, if data for pre-training and fine-tuning are the same, it is possible to train in one go, instead of dividing in two stages. Or you can skip pre-training and use only multilayer neural network, or, on the other hand, use only RBM without the neural network. At the same time we have access to all internal parameters. The package is intended for advanced users. Further, we will analyze divided processes: pre-training and fine-tuning.
3.1.2. Building the DBN model. Parameters.
We will describe the process of building, training and testing the DBN model.
1. We create the deep architecture object named ‘Darch’ using the constructor with necessary parameters
newDArch(layers, batchSize, ff=FALSE, logLevel=INFO, genWeightFunc=generateWeights),
where:
- layers: array indicating the number of layers and neurons in each layer. For example: layers = c(5,10,10,2) – an input layer with 5 neurons (visible), two hidden layers with 10 neurons each, and one output layer with 2 outputs.
- BatchSize: size of the mini-sample during training.
- ff: indicates whether the ff format should be used for weights, deviations and exits. The ff format is applied for saving large volumes of data with compression.
- LogLevel: level of logging and output when performing this function.
- GenWeightFunction: function for generating the matrix of RBM weights. There is an opportunity to use the user’s activation function.
The created darch-object contains (layers – 1) RBM combined into the accumulating network that will be used for pre-training the neural network. Two attributes fineTuneFunction and executeFunction contain functions for fine-tuning (backpropagation by default) and for execution (runDarch by default). Training the neural network is performed with two training functions: preTrainDArch() and fineTuneDArch(). The first function trains the RBM network without a supervisor using a contrastive divergence method. The second function uses a function indicated in the fineTuneFunction attribute for a fine-tuning of neural network. After neural network performance, outputs of every layer can be found in the executeOutputs attribute or only output layer in the executeOutput attribute.
2. Function of pre-training the darch-object
preTrainDArch(darch, dataSet, numEpoch = 1, numCD = 1, …, trainOutputLayer = F),where:
- darch: instance of the ‘Darch’ class;
- dataSet: data set for training;
- numEpoch: number of training epochs;
- numCD : number of sampling iterations. Normally, one is sufficient;
- … : additional parameters that can be transferred to the trainRBM function;
- trainOutputLayer: logical value that shows whether the output layer of RBM should be trained.
The function performs the trainRBM() training function for every RBM, by copying after training weights and biases to the relevant neural network layers of the darch-object.
3. Fine-tune function of the darch-object
fineTuneDArch(darch, dataSet, dataSetValid = NULL, numEpochs = 1, bootstrap = T,
isBin = FALSE, isClass = TRUE, stopErr = -Inf, stopClassErr = 101,
stopValidErr = -Inf, stopValidClassErr = 101, ...),
where:
- darch: sample of the ‘Darch’ class;
- dataSet: set of data for training (can be used for validation) and testing;
- dataSetValid : set of data used for validation;
- numxEpoch: number of training epochs;
- bootstrap: logical, is it needed to apply bootstrap when creating validation data;
- isBin:indicates if output data should be interpreted as logical values. By default — FALSE. If TRUE, every value above 0.5 is interpreted as 1, and below — as 0.
- isClass : indicates if the network is trained for classification. If TRUE, then statistics for classifications will be determined. TRUE by default.
- stopErr : criterion for stopping the training of neural network due to error occurred during training. -Inf by default;
- stopClassErr : criterion for stopping the training of neural network due to classification error occurred during training. 101 by default;
- stopValidErr : criterion for stopping the neural network due to error in validation data. -Inf by default;
- stopValidClassErr : criterion for stopping the neural network due to classification error occurred during validation. 101 by default;
- … : additional parameters that can be passed to the training function.
The function trains the network with a function saved in the fineTuneFunction attribute of the darch-object. Input data (trainData, validData, testData) and classes that belong to them (targetData, validTargets, testTargets) can be transferred as dataSet or ff-matrix. Data and classes for validation and testing are not obligatory. If they are provided, then neural network will be performed with these sets of data, and statistics will be calculated. The isBin attribute indicates if output data should be interpreted as binary. If isBin = TRUE, every output value above 0.5 is interpreted as 1, otherwise — as 0. Also, we can set a stopping criterion for the training based on error (stopErr, stopValidErr) or correct classification (stopClassErr, stopValidClassErr) on training or validation sets.
All function parameters have default values. However, other values are also available. So, for example:
Function of activating neurons — sigmoidUnitDerivative, linearUnitDerivative, softmaxUnitDerivative, tanSigmoidUnitDerivative are available. sigmoidUnitDerivative is used by default.
Functions of the neural network’s fine-tune — backpropagation by default, resilient-propagation rpropagation is also available in four variations (“Rprop+”, “Rprop-“, “iRprop+”, “iRprop-“) and minimizeClassifier (this function is trained by the Darch network classifier using the nonlinear conjugate gradient method). For the last two algorithms and for those who have a deep knowledge of the subject, a separate implementation of the neural network’s fine-tune with a configuration of their multiple parameters is provided. For example:
rpropagation(darch, trainData, targetData, method="iRprop+",
decFact=0.5, incFact=1.2, weightDecay=0, initDelta=0.0125,
minDelta=0.000001, maxDelta=50, ...),
where:
- darch – darch-object for training;
- trainData – input data set for training;
- targetData – expected output for the training set;
- method – training method. “iRprop+” by default. “Rprop+”, “Rprop-“, “iRprop-” are possible;
- decFact – decreasing factor for training. 0.5 by default;
- incFact – increasing factor for training. 1.2 by default;
- weightDecay – decreasing weight at the training. 0 by default;
- initDelta – initialization value at the update. 0.0125 by default;
- minDelta – minimum border for the step size. 0.000001 by default;
- maxDelta – upper border for the step size. 50 by default.
The function returns the darch-object with the trained neural network.
3.2. Formation of training and testing samples.
We have already formed the initial sample of data. Now, we need to divide it into training, validating and testing samples. The ratio by default is 2/3. Various packages have many functions that are used to divide samples. I use rminer::holdout() that calculates indexes for breaking down the initial sample into training and testing samples.
holdout(y, ratio = 2/3, internalsplit = FALSE, mode = "stratified", iter = 1,
seed = NULL, window=10, increment=1),
where:
- y – desired target variable, numeric vector or factor, in this case, the stratified separation is applied (i.e. proportions between the classes are the same for all parts);
- ratio – ratio of separation (in percentage — the size of the training sample is established; or in the total number of samples — the size of the testing sample is established);
- internalsplit – if TRUE, then training data is once again separated into training and validation samples. The same ratio is applied for the internal separation;
- mode – sampling mode. Options available:
- stratified – stratified random division (if у factor; otherwise standard random division);
- random – standard random division;
- order – static mode, when first examples are used for training, and the remaining ones — for testing (widely applied for time series);
- rolling – rolling window more commonly known as a sliding window (widely applied at the prediction of stock and financial markets), similarly order, except that window refers to window size, iter — rolling iteration and increment — number of samples the window slides forward at every iteration. The size of the training sample for every iteration is fixed with window, while the testing sample is equivalent to ratio, except for the last iteration (where it could be less).
- incremental – incremental mode of re-training, also known as an increasing window, same as order, except that window is an initial window size, iter — incremental iterations and increment — number of examples added at every iteration. The size of the training sample grows (+increment) at every iteration, whereas the size of the testing set is equivalent to ratio, except for the last iteration, where it can be smaller.
- iter – number of iterations of the incremental mode of re-training (used only when mode = “rolling” or “incremental”, iter is usually set in a loop).
- seed – if NULL, then random seed is used, otherwise seed is fixed (further calculations will always have the same result returned);
- window – size of training window (if mode = “rolling”) or the initial size of training window (if mode = “incremental”);
- increment – number of examples added to the training window at every iteration (if mode=”incremental” or mode=”rolling”).
3.2.1. Balancing classes and pre-processing.
We will write a function that will align (if required) the number of classes in the sample towards the higher number, divide the sample into training and testing samples, perform pre-processing (normalization, if necessary) and return the list with relevant samples — train, test. To achieve balancing, we are going to use the caret::upSample() function that adds samples randomly taken with replacement, making the class distribution equal. I must say that not all researchers find it necessary to balance classes. But, as already known, practice is a criterion of truth, and the results of my multiple experiments show that balanced samples always show better results in training. Although, it doesn’t stop us to experiment on our own.
For pre-processing we will use the caret::preProcess() function. Parameters of preprocessing will be saved in the prepr variable. Since we have already considered and applied them in previous articles, I will not provide any further description here.
#---7---------------------------------------------------- prepareTrain <- function(x , y, rati, mod = "stratified", balance = F, norm, meth) { require(magrittr) require(dplyr) t <- rminer::holdout(y = y, ratio = rati, mode = mod) train <- cbind(x[t$tr, ], y = y[t$tr]) if(balance){ train <- caret::upSample(x = train[ ,best], y = train$y, list = F)%>% tbl_df train <- cbind(train[ ,best], select(train, y = Class)) } test <- cbind(x[t$ts, ], y = y[t$ts]) if (norm) { prepr <<- caret::preProcess(train[ ,best], method = meth) train = predict(prepr, train[ ,best])%>% cbind(., y = train$y) test = predict(prepr, test[ ,best] %>% cbind(., y = test$y)) } DT <- list(train = train, test = test) return(DT) }
One comment regarding pre-processing: input variables will be normalized into the range of (-1, 1).
3.2.2. Coding the target variable
When solving classification tasks, the target variable is a factor with several levels (classes). In a model it is set as a vector (column), that consists of the subsequent target states. For example, y = с(“1”, “1”, “2”, “3”, “1”). In order to train the neural network, the target variable must be coded (transformed) into the matrix with the number of columns equal to the number of classes. In every row of this matrix, only one column may contain 1. Such transformation along with using the softmax() activation function in the output layer, allows to obtain probabilities of states of the predicted target variable in every class. The classvec2classmat() function will be used for coding. This in not the only or the best method for coding the target variable, but we will use it because it is simple. Inverse transformation (decoding) of predicted values of the target variable is achieved through different methods that we are going to cover soon.
3.3. Training the model
3.3.1. Pre-training
As mentioned above, first, we create the deep architecture object named DArch, that includes the required number of RBM with parameters of preliminary training by default, and the neural network initiated with random weights and neuron activation function set by default. At the object creation stage, the pre-training parameters can be changed, if necessary. Afterwards, the RBM network will be pre-trained without a supervisor by sending the training sample (without target variable) to the output. After it is complete, we get DАrch where weights and biases obtained during RBM training are transferred to the neural network. We should set in advance the distribution of hidden neurons in layers in a form of vector (for example):
L<- c( 14, 50, 50, 2)
Number of neurons in the input layer equals the number of input variables. Two hidden layers will contain 50 neurons each, the output layer will have two. Let me explain the last bit. If a target variable (factor) has two levels (classes), then, in fact, one output is sufficient. But converting vector into the matrix with two columns, each of them corresponding to one class, allows us to apply the softmaxactivation function, that operates well in the classification tasks, in the output layer. Furthermore, outputs in the form of the class probabilities give us additional opportunities in the subsequent analysis of results. This subject will be covered shortly.
The number of epochs when training is set experimentally, normally within range of 10-50.
The number of sampling iteration will stay by default, but this parameter can be increased if you wish to experiment. It will be defined in a separate function:
#----8-------------------------------------------------------------- pretrainDBN <- function(L, Bs, dS, nE, nCD, InM = 0.5, FinM = 0.9) { require(darch) # create object DArch dbn <- newDArch(layers = L, batchSize = Bs, logLevel = 5) # set initial moment setInitialMomentum(dbn) <- InM # set final moment setFinalMomentum(dbn) <- FinM # set time of switching moments from initial to final setMomentumSwitch(dbn) <- round(0.8 * nE) dbn <- preTrainDArch(dbn, dataSet = dS, numEpoch = nE, numCD = nCD, trainOutputLayer = T) return(dbn) }
3.3.2. Fine-tuning
As discussed previously, the package offers backpropagation(), rpropagation(), minimizeClassifier(), minimizeAutoencoder() for fine-tuning. The last two won’t be considered, since they are not sufficiently documented in the package, and there are no examples of how to apply them. These functions in my experiments didn’t show good results.
I would also like to add something about package updates. When I started writing this article, the current version was 0.9, and by the moment it was finished, a new 0.10 version containing multiple changes was released. All calculations had to be redone. Based on the results of short tests, I can tell that the operation speed has considerably increased, unlike the results’ quality (which is more a fault of a user, then the package).
Let’s consider two first functions. The first (backpropagation) is set by default in the DАrch object and uses the training neural network parameters provided here. The second function (rpropagation) also has default parameters and four training methods (described above) that are “iRprop+” by default. You can certainly change both parameters and the training method. It is easy to apply these functions: change the fine-tune function in FineTuneDarch()
setFineTuneFunction(dbn) <- rpropagation
In addition to fine-tuning settings, we must set (if necessary) the function of activating neurons in every layer. We know that sigmoidUnit is set in all layers by default. It is available in the package sigmoidUnitDerivative, linearUnitDerivative, tanSigmoidUnitDerivative, softmaxUnitDerivative . The fine-tune will be defined with a separate function with the ability to choose the fine-tune function. We will collect possible functions of activation in a separate list:
actFun <- list(sig = sigmoidUnitDerivative, tnh = tanSigmoidUnitDerivative, lin = linearUnitDerivative, soft = softmaxUnitDerivative)
We will write a fine-tune function that will train and generate two neural networks: first — trained using the backpropagation function, second — with rpropagation:
#-----9----------------------------------------- fineMod <- function(variant=1, dbnin, dS, hd = 0.5, id = 0.2, act = c(2,1), nE = 10) { setDropoutOneMaskPerEpoch(dbnin) <- FALSE setDropoutHiddenLayers(dbnin) <- hd setDropoutInputLayer(dbnin) <- id layers <<- getLayers(dbnin) stopifnot(length(layers)==length(act)) if(variant < 0 || variant >2) {variant = 1} for(i in 1:length(layers)){ fun <- actFun %>% extract2(act[i]) layers[[i]][[2]] <- fun } setLayers(dbnin) <- layers if(variant == 1 || variant == 2){ # backpropagation if(variant == 2){# rpropagation #setDropoutHiddenLayers(dbnin) <- 0.0 setFineTuneFunction(dbnin) <- rpropagation } mod = fineTuneDArch(darch = dbnin, dataSet = dS, numEpochs = nE, bootstrap = T) return(mod) } }
Some clarifications about formal parameters of the function.
- variant – selection of fine-tune function (1- backpropagation, 2- rpropagation).
- dbnin – model of receipt resulted from pre-training.
- dS – data set for fine-tune (dataSet).
- hd – coefficient of sampling (hiddenDropout) in hidden layers of neural network.
- id – coefficient of sampling (inputDropout) in input layer of neural network.
- act – vector with indication of function of neuron activation in every layer of neural network. The length of vector is one unit shorter than the number of layers.
- nE – number of training epochs.
dataSet — a new variable that appeared in this version. I don’t really understand the reason behind its appearance. Normally, the language has two ways of transferring variables to a model — using a pair (x, y) or a formula (y~., data). The introduction of this variable doesn’t improve the quality, but confuses the users instead. However, the author may have his reasons that are unknown to me.
3.4. Testing the model. Мetrics.
Testing the trained model is performed on testing samples. It must be considered that we will calculate two metrics: formal Accuracy and qualitative K. The relevant information will be provided below. For this purpose, we will need two different samples of data, and I will explain to you why. To calculate Accuracy we need values of the target variable, and the ZigZag, as we remember from before, most frequently is not defined on the last bars. Therefore, the testing sample for calculating Accuracy we will determine with the prepareTrain() function, and for qualitative indicators we will use the following function
#---10------------------------------------------- prepareTest <- function(n, z, norm, len = 501) { x <- In(p = n ) %>% na.omit %>% extract( ,best) %>% tail(., len) CO <- price[ ,"CO"] %>% tail(., len) if (norm) { x <- predict(prepr,x) } dt <- cbind(x = x, CO = CO) %>% as.data.frame() return(dt) }
The models will be tested on the last 500 bars of the history.
For actual testing, testAcc() and testBal() will be applied.
#---11----- testAcc <- function(obj, typ = "bin"){ x.ts <- DT$test[ ,best] %>% as.matrix() y.ts <- DT$test$y %>% as.integer() %>% subtract(1) out <- predict(obj, newdata = x.ts, type = typ) if (soft){out <- max.col(out)-1} else {out %<>% as.vector()} acc <- length(y.ts[y.ts == out])/length(y.ts) %>% round(., digits = 4) return(list(Acc = acc, y.ts = y.ts, y = out)) } #---12----- testBal <- function(obj, typ = "bin") { require(fTrading) x <- DT.test[ ,best] CO <- DT.test$CO out <- predict(obj, newdata = x, type = typ) if(soft){out <- max.col(out)-1} else {out %<>% as.vector()} sig <- ifelse(out == 0, -1, 1) sig1 <- Hmisc::Lag(sig) %>% na.omit bal <- cumsum(sig1 * tail(CO, length(sig1))) K <- tail(bal, 1)/length(bal) * 10 ^ Dig Kmax <- max(bal)/which.max(bal) * 10 ^ Dig dd <- maxDrawDown(bal) return(list(sig = sig, bal = bal, K = K, Kmax = Kmax, dd = dd)) }
The first function returns Acc and the target variable values (real or predicted) for a possible further analysis. The second function returns the predicted signals sig for the EA, the balance obtained based on these signals (bal), quality coefficient (К), maximum value of this coefficient on the tested area (Kmax) and the maximum drawdown (dd) in the same area.
When calculating the balance, it is important to remember that the last predicted signal refers to the future bar that hasn’t been formed yet, therefore, it should be deleted at calculations. We have done it by moving the sig vector by one bar to the right.
3.4.1. Decoding predictions.
The obtained result can be decoded (converted from matrix to vector) using the “WTA” method. The class equals the column number with a maximum value of probability, and the value threshold of this probability can be set, below which the class is not determined.
out <- classmat2classvec(out, threshold = 0.5) or out <- max.col(out)-1
If the threshold is set as 0.5, and the biggest probability in the columns is below this threshold, we will obtain an additional class (“not defined”). It should be taken into consideration when calculating metrics like Accuracy.
3.4.2. Improving the prediction results
Is it possible to improve the prediction result after it is received? There are three possible ways that can be applied.
- Calibration
Calibration is a calculation of the possibility ranges that give the most accurate compatibility with real data. For this purpose, there is a special function in the CORElearn package:
CORElearn::calibrate(correctClass, predictedProb, class1 = 1, method = c("isoReg", "binIsoReg", "binning", "mdlMerge"), weight=NULL, noBins=10, assumeProbabilities=FALSE)
where:
- correctClass — vector with correct labels of classes for problem classification;
- predictedProb — vector with the predicted class 1 (probability) of the same length as correctClass;
- method — one out of the following (“isoReg”, “binIsoReg”, “binning”, “mdlMerge”). For further information please read the package description;
- weight — vector (if indicated) must be the same length as correctClass, and provide weights for each observation, otherwise weights of all observations equal 1 by default;
- noBins — parameter value depends on method and determines the desired or initial number of channels;
- assumeProbabilities — logical, if TRUE, then value in predictedProb is expected in the range [0, 1], i. e. as a possibility evaluation, otherwise the algorithm can be used as a simple isotonic regression.
This method is applied for a target variable with two levels set by a vector.
- Smoothing prediction results with the Markov chain model
This is a vast and complex subject that deserves a separate article, therefore I won’t go deep into theory, and provide the most basic information.
Markov’s process — is a random process with a following feature: for any point in time t0, probability of any state of the system in the future depends only on its state in the present and doesn’t depend on when and how the system reaches this state.
Classification of random Markov’s processes:
- with discrete states and discrete time (Markov chain);
- with continuous states and discrete time (Markov consistency);
- with discrete states and continuous time (continuous Markov chain);
- with continuous state and continuous time.
Only Markov processes with discrete states S1, S2, …, Sn. are considered here further.
Markov chain — random Markov process with discrete states and discrete time.
Moments t1, t2, … when the S system can change its state are considered as subsequent steps of the process. It is not the t time, but the step number 1,2,…,k,…that is used as an argument that the process depends on.
Random process is characterized by a sequence of states S(0), S(1), S(2), …, S(k), …, where S(0) is the initial state (before the first step); S(1) — state after the first step; S(k) — state of the system after the k-number step.
Probabilities of the Markov chain states are Pi(k) probabilities that after the k-number step (and before (k + 1)-step) the S system will be in the Si(i = 1 , 2 , …, n) state.
Initial distribution of the Markov chain probabilities — distribution of the probabilities of states in the beginning of the process.
Probability of transition (transition probability) on the k-number step from the Si state to the Sj state — conditional probability that the S system after the k-number step will appear in the Sj state, on condition that it was in the Si state before that (after k—1 step).
Uniform Markov chain — Markov chain where transition probabilities don’t depend on the step number (from time), but on between which states the transition takes place.
Transitional probabilities of a uniform Markov chain Рij form a square matrix sized n х n. It has the following features:
- Each row describes the selected state of the system, and its elements — probabilities of all possible transitions in one step from the selected (from i-number) state.
- Elements of columns — probabilities of all possible transitions in one step to the set (j) state.
- The total of probabilities of each row equals one.
- On the main diagonal — Рij probabilities that the system won’t exit from the Si state, and will remain there.
Markov process can be observed and hidden. Hidden Markov Model (HMM) consists of the pair of discrete stochastic processes {St} and {Xt}. The observed process {Xt} is linked with an unobserved (hidden) Markov process of states {St} through so-called conditional distributions.
Strictly speaking, the observed Markov process of states (classes) of our target time series is not uniform. Clearly, the probability of transition from one state to another depends on the time spent in the current state. It means that during the first steps after changing the state, the probability that it will change is low and grows with the increase of time spent in this state. These models are called semi-Markov’s (HSMM). We won’t go deep into analyzing them.
But the idea is the following: based on the discrete order of ideal signals (target) obtained from the ZigZag, we will find the parameters of НММ. Then, having the signals predicted by the neural network, we will smooth them using НММ.
What does it give us? Normally, in the neural network prediction there are so-called “emissions”, areas of changing the state that is 1-2 bars long. We know that a target variable doesn’t have such small lengths. By applying the model obtained on the target variable to the predicted order, we can bring it to more probable transitions.
We will use the “mhsmm” package designed for calculating hidden Markov and semi-Markov models for these calculations. We will use the smooth.discrete() function, that simply smooths the time series of discrete values.
obj <- smooth.discrete(y)
Smooth order of states obtained in the end by default — as a more likely order of states obtained using the Viterbi algorithm (so called global decoding). There is also an option to use other method — smoothed, where individual most probable states will be identified (so-called local decoding).
A standard method is applied to smooth new time series
sm.y <- predict(obj, x = new.y)
- Correcting predicted signals on the theoretical balance curve
The concept is the following. Having the balance line, we can calculate its deviation from the average one. Using these deviations we will calculate correction signals. In moments when deviations go minus, they either disable the performance of predicted signals, or make them inverse. The idea is generally good, but there is one disadvantage. The zero bar has a predicted signal, but it doesn’t have a balance value and, therefore, a correction signal. There are two ways to solve this issue: through classification — to predict correction signal based on existing correction signals and deviations; through regression — using the existing deviations on the formed bars to predict deviations on the new bar and to identify the correction signal based on it. There is an easier solution, where you can take the correction signal for a new bar on the basis of the one that is already formed.
Since the above mentioned methods are already known to us and have been tested, we will try to implement opportunities of the Markov chains.The “markovchain” package that appeared recently has a range of functions that allow to determine the parameters of the hidden Markov model and to project future states by several future bars through the observed discrete process. The idea was taken from this article.
3.4.3. Metrics
To evaluate the quality of model prediction, the whole range of metrics (Accuracy, AUC, ROC and other) is applied. In the previous article I have mentioned that formal metrics can’t define quality in our case. The Expert Advisor’s goal is to get the maximum profit with an acceptable drawdown. For this purpose, K quality indicator was introduced, and it shows average profit in points for one bar on the fixed history segment with N length. It is calculated through dividing the cumulative Return(sig, N) by the length of the N segment. Accuracy will be calculated only indicatively.
Finally, we will perform calculations and obtain testing results:
- Output data. We already have the price[] matrix, obtained as a result of performing the price.OHLC() function. It contains quotes, average price and body of the bars. All output data can be obtained by downloading the “icon” that appears in the attachment to Rstudio.
# Find constanta n = 34; z = 37; cut = 0.9; soft = TRUE. # Find preprocessing method method = c("center", "scale","spatialSign") # form the initial set of data data.f <- form.data(n = n, z = z) # find the set of important predictors best <- prepareBest(n = n, z = z, cut = cut, norm = T, method) # Calculations take about 3 minutes on the 2-core processor. You can skip this stage if you like, # and use the whole set of predictors in the future. Therefore, comment the previous line and # uncomment two lowest lines. # data.f <- form.data(n = n, z = z) # best <- colnames(data.f) %>% head(., ncol(data.f) - 1) # Prepare the set for training neural network DT <- prepareTrain(x = data.f[ , best], y = data.f$y, balance = TRUE, rati = 501, mod = "stratified", norm = TRUE, meth = method) # Download required libraries require(darch) require(foreach) # Identify available functions for activation actFun <- list(sig = sigmoidUnitDerivative, tnh = tanSigmoidUnitDerivative, lin = linearUnitDerivative, soft = softmaxUnitDerivative) # Convert the target variable if (soft) { y <- DT$train$y %>% classvec2classmat()} # into matrix if (!soft) {y = DT$train$y %>% as.integer() %>% subtract(1)} # to vector with values [0, 1] # create dataSet for training dataSet <- createDataSet( data = DT$train[ ,best] %>% as.matrix(), targets = y , scale = F ) # Identify constants for neural network # Number of neurones in the input layer (equals the amount of predictors) nIn <- ncol(dataSet@data) # Number of neurones in the output layer nOut <- ncol(dataSet@targets) # Vector with a number of neurones in every layer of neural network # If you use another structure of neural network, this vector should be rewritten Layers = c(nIn, 2 * nIn , nOut) # Other data related to training Bath = 50 nEp = 100 ncd = 1 # Pre-training of neural network preMod <- pretrainDBN(Layers, Bath, dataSet, nEp, ncd) # Additional parameters for fine-tune Hid = 0.5; Ind = 0.2; nEp = 10 # Train two models, one with backpropagation, other with rpropagation # We only do this to compare results model <- foreach(i = 1:2, .packages = "darch") %do% { dbn <- preMod if (!soft) {act = c(2, 1)} if (soft) {act = c(2, 4)} fineMod(variant = i, dbnin = dbn, hd = Hid, id = Ind, dS = dataSet, act = act, nE = nEp) } # Test to get Accuracy resAcc <- foreach(i = 1:2, .packages = "darch") %do% { testAcc(model[[i]]) } # Prepare sample of data to test on quality coefficient DT.test <- prepareTest(n = n, z = z, T) # Test resBal <- foreach(i = 1:2, .packages = "darch") %do% { testBal(model[[i]]) }
Let’s see the result:
> resAcc[[1]]$Acc [1] 0.5728543 > resAcc[[2]]$Acc [1] 0.5728543
It is equally bad for both models.
As for the quality coefficient:
> resBal[[1]]$K [1] 5.8 > resBal[[1]]$Kmax [1] 20.33673 > resBal[[2]]$Kmax [1] 20.33673 > resBal[[2]]$K [1] 5.8
It shows the same good performance. However, a large drawdown is somehow alarming:
> resBal[[1]]$dd$maxdrawdown [1] 0.02767
We will try to correct the drawdown with a correction signal obtained from the below calculation:
bal <- resBal[[1]]$bal # signal on the last 500 bars sig <- resBal[[1]]$sig[1:500] # average from the balance line ma <- pracma::movavg(bal,16, "t") # momentum from the average roc <- TTR::momentum(ma, 3)%>% na.omit # balance line deviation from the average dbal <- (bal - ma) %>% tail(., length(roc)) # summarize two vectors dbr <- (roc + dbal) %>% as.matrix() # calculate correction signal sig.cor <- ifelse(dbr > 0, 1, -1) # sign(dbr) gives the same result # resulting signal S <- sig.cor * tail(sig, length(sig.cor)) # balance on resulting signal Bal <- cumsum(S * (price[ ,"CO"]%>% tail(.,length(S)))) # quality coefficient on the corrected signal Kk <- tail(Bal, 1)/length(Bal) * 10 ^ Dig > Kk [1] 28.30382
The shown quality result on the corrected signal is very good. Let’s see how the lines dbal, roc and dbr used for calculating the correction signal appear on the line chart.
matplot(cbind(dbr, dbal, roc), t="l", col=c(1,2,4), lwd=c(2,1,1)) abline(h=0, col=2) grid()
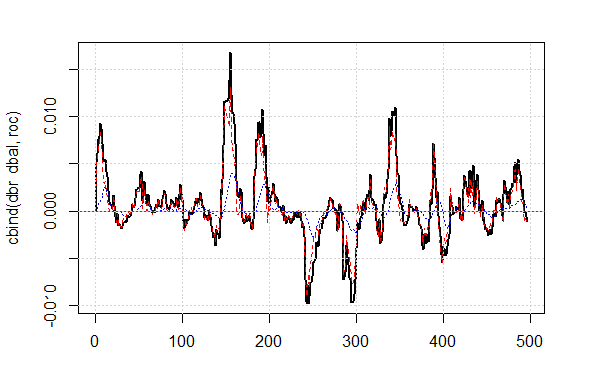
Fig.4 Balance line deviation from the average
Balance line before and after the signal correction is shown in fig. 5.
plot(c(NA,NA,NA,Bal), t="l") lines(bal, col= 2) lines(ma, col= 4)
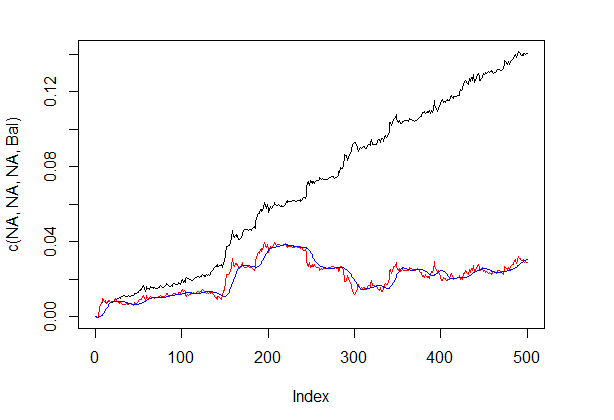
Fig.5 Balance line before and after the signal correction
So, we have the signal’s value predicted by the neural network on the zero bar, but don’t have a correction value. We want to use the hidden Markov model for predicting this signal. Based on the observed states of the correction signal we will identify the model’s parameters using values of few last states, predict the state at one bar ahead. First, we will write the correct() function, that will calculate the correction signal based on the predicted one, resulting signal and its quality indicators. In other words, we will compactly write down calculations performed previously.
I wish to clarify: the “signal” in the article is a sequence of integer -1 and 1. The “state” is a sequence of integers 1 and 2 corresponding to these signals. For mutual conversions we will use the functions:
#---13---------------------------------- sig2stat <- function(x) {x %>% as.factor %>% as.numeric} stat2sig <- function(x) ifelse(x==1, -1, 1) #----14--correct----------------------------------- correct <- function(sig){ sig <- Hmisc::Lag(sig) %>% na.omit bal <- cumsum(sig * (price[ ,6] %>% tail(.,length(sig)))) ma <- pracma::movavg(bal, 16, "t") roc <- TTR::momentum(ma, 3)%>% na.omit dbal <- (bal - ma) %>% tail(., length(roc)) dbr <- (roc + dbal) %>% as.matrix() sig.cor <- sign(dbr) S <- sig.cor * tail(sig, length(sig.cor)) bal <- cumsum(S * (price[ ,6]%>% tail(.,length(S)))) K <- tail(bal, 1)/length(bal) * 10 ^ Dig Kmax <- max(bal)/which.max(bal) * 10 ^ Dig dd <- fTrading::maxDrawDown(bal) corr <<- list(sig.c = sig.cor, sig.res = S, bal = bal, Kmax = Kmax, K = K, dd = dd) return(corr) }
In order to obtain the signal vector with prediction of 1 bar ahead, we will use the “markovchain” package and the pred.sig() function.
#---15---markovchain---------------------------------- pred.sig <- function(sig, prev.bar = 10, nahead = 1){ require(markovchain) # Transform the observed correction signals into states stat <- sig2stat(sig) # Calculate model parameters # if there is no model in the environment if(!exists('MCbsp')){ MCbsp <<- markovchainFit(data = stat, method = "bootstrap", nboot = 10L, name="Bootstrap MС") } # Set necessary constants newData <- tail(stat, prev.bar) pr <- predict(object = MCbsp$estimate, newdata = newData, n.ahead = nahead) # attach the predicted signal to input signal sig.pr <- c(sig, stat2sig(pr)) return(sig.pr = sig.pr) }
Now, we will write down the resulting signal calculation for the Expert Advisor to perform compactly:
sig <- resBal[[1]]$sig sig.cor <- correct(sig) sig.c <- sig.cor$sig.c pr.sig.cor <- pred.sig(sig.c) sig.pr <- pr.sig.cor$sig.pr # Resulting vector of signals for Expert Advisor S <- sig.pr * tail(sig, length(sig.pr))
- Smoothing the predicted signal.
We will write a function that will smooth the discrete signal using the model of the hidden Markov chain. For this purpose, we will use the “mhsmm” package.
#---16---smooth------------------------------------
smoooth <- function(sig){
# smooth predicted signal
# define parameters of hidden Markov model
# if there is no model in the environment yet
require(mhsmm)
if(!exists('obj.sm')){
obj.sm <<- sig2stat(sig)%>% smooth.discrete()
}
# smooth the signal with the obtained model
sig.s <- predict(obj.sm, x = sig2stat(sig))%>%
extract2(1)%>% stat2sig()
# calculate balance with smoothed signal
sig.s1 <- Hmisc::Lag(sig.s) %>% na.omit
bal <- cumsum(sig.s1 * (price[ ,6]%>% tail(.,length(sig.s1))))
K <- tail(bal, 1)/length(bal) * 10 ^ Dig
Kmax <- max(bal)/which.max(bal) * 10 ^ Dig
dd <- fTrading::maxDrawDown(bal)
return(list(sig = sig.s, bal = bal, Kmax = Kmax, K = K, dd = dd))
}
We will calculate and compare the balance based on predicted and smoothed signals.
sig <- resBal[[1]]$sig sig.sm <- smoooth(sig) plot(sig.sm$bal, t="l") lines(resBal[[1]]$bal, col=2)
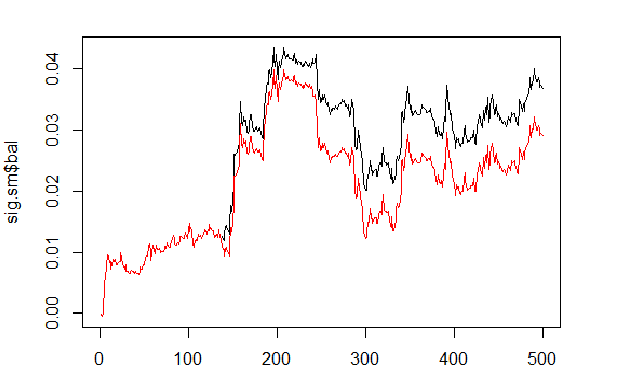
Fig.6 Balance based on smoothed and predicted signals
As we can see, the quality has slightly improved, but the drawdown still remains. We won’t use this method in our Expert Advisor.
sig.sm$dd $maxdrawdown [1] 0.02335 $from [1] 208 $to [1] 300
4. Structure of the EA algorithm
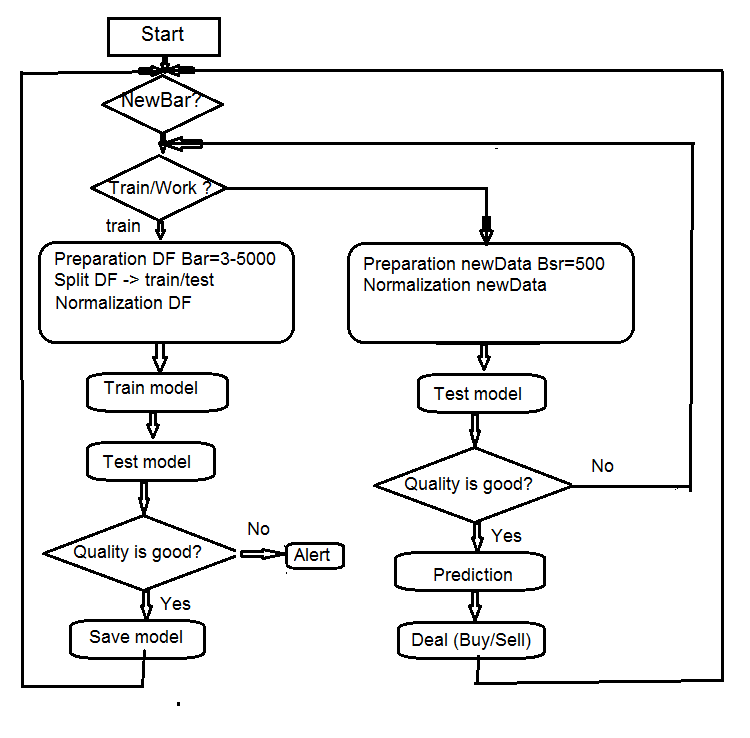
Fig.7 Structure of the EA algorithm
4.1. Description of the Expert Advisor’s operation
Since the Expert Advisor operates in two streams (mql and Rterm), we will describe the process of their interaction. We will discuss the operations performed in each stream separately.
4.1.1 MQL
After placing the Expert Advisor on the chart:
in the init() function
- we check the terminal’s settings (DLL availability, permission to trade);
- set the timer;
- launch Rterm;
- calculate and transfer constants required for work to the R-process environment;
- check if Rterm works, if not – alert;
- exit from init().
In the deinit() function
- we stop the timer;
- delete graphic objects;
- stop the Rterm.
In the onTimer()function
- check if Rterm is working;
- if Rterm is not occupied and the new bar is (LastTime != Time[0]):
- set the depth of history depending on if this is a first launch of the Expert Advisor;
- form four vectors of quotes (Open, High, Low, Close) and transfer them to Rterm;
- launch the script and leave without receiving the results of its performance;
- set the get_sig = true flag;
- set LastTime= Time[0].
- Otherwise, if Rterm works, is not occupied and the flag is get_sig = true:
- identify length of the sig vector that we should receive from Rterm;
- adjust the size of the receiving vector to the size of the source. When failing to comply, Rprocess will drop;
- obtain signals’ order (vector);
- determine which operation has to be performed (BUY, SELL, Nothing) using the last signal;
- if we obtain the real signal, not ERR, we reset the flag get_sig=false.
- the rest is standard:
- CheckForClose()
- CheckForOpen()
Our expert in this part is a “Performer” that carries out orders obtained from its part that can “think”, it sends orders, tracks the state of open positions and possible errors when opening them, and performs many other functions of a standard Expert Advisor.
4.1.2 Rterm
The operating script consists of two parts. One part is executed at the first entry, second — at the following ones.
- if first:
- upload (if required) necessary libraries from the depositary on the Internet, and install them to the environment of Rterm;
- define necessary functions;
- create the quote matrix;
- prepare the sample of data for training and testing the model;
- create and train the model;
- test the model;
- calculate signals for performance;
- check the quality of prediction. If it is above or equals the set minimum — proceed. Otherwise — send alert.
- if !first:
- prepare the sample of data for testing and prediction;
- test the model on new data;
- calculate signals for performance;
- check the quality of prediction. If it exceeds or equals the set minimum — we proceed. Otherwise — we set first = TRUE, i.e. we request to re-train the model.
4.2. Self-control and Self-training
The quality control of predicting signals with a model is performed using the К coefficient. There are two ways to identify limits of the acceptable quality. First — to set the maximum fall of the coefficient in relation to its maximum value. If К < Kmax * 0.8, then we should re-train or stop the Expert Advisor from performing signals. Second — to set the minimum value of К, that after being reached requires the same actions. We will use the second method in the Expert Advisor.
5. Installation and launching
There are two Expert Advisors attached to this article: e_DNSAE.mq4 and e_DNRBM.mq4. Both of them use the same samples of data and almost the same set of functions. The difference lies in the deep network model used. The first EA uses DN, initiated SAE and the “deepnet” package. The package description can be found in the previous article on deep neural networks. The second EA uses DN, initiated RBM and the “darch” package.
Standard distribution applies:
- *.mq4 in the ~/MQL4/Expert folder
- *.mqh in the ~/MQL4/Include folder
- *.dll in the ~/MQL4/Libraries folder
- *.r in the C:/RData folder
We correct the path to the set R language and scripts (in both mq4: #define and *.r: source() ).
When the Expert Advisor is launched for the first time, it will download necessary libraries from the repository and set them in the Rterm environment. You can also install them in advance according to the list attached.
Normally, the R process “drops” specifically due to the absence of necessary libraries, wrongly indicated paths to scripts, and only lastly, because of the script syntax errors.
The session’s screenshot with the initial data is attached separately, and you can open it with Rstudio to check that all functions are working, as well as conduct the experiments.
6. Ways and methods of improving qualitative indicators.
There are few ways to improve qualitative indicators.
- evaluation and selection of predictors — apply genetic algorithm of optimization (GA).
- determine optimal parameters of predictors and target variable — GA.
- determine optimal parameters of a neural network — GA.
Taking these measures helps to considerably improve qualitative indicators.
Conclusion
Experiments with the “darch” package have shown the following results.
- The deep neural network, initiated RBM are trained worse than with SAE. This is hardly news for us.
- The network is trained quickly.
- The package has a big potential in improving the quality of prediction, by providing access to almost all internal parameters of the model.
- The package allows to use only a neural network or RBM with a very wide set of parameters in relation to other, standard ones.
- The package is constantly evolving, and the developer promises to introduce additional features in the next release.
- R language integration with the МТ4/МТ5 terminals, as promised by the developers, will give traders an opportunity to use the newest algorithms without any additional DLL.
Awesome https://shorturl.fm/5JO3e
Very good partnership https://shorturl.fm/68Y8V
Very good https://shorturl.fm/TbTre
Cool partnership https://shorturl.fm/FIJkD
Cool partnership https://shorturl.fm/XIZGD
Super https://shorturl.fm/6539m
Best partnership https://shorturl.fm/A5ni8
Good partner program https://shorturl.fm/m8ueY
Best partnership https://shorturl.fm/A5ni8
Good partner program https://shorturl.fm/m8ueY
https://shorturl.fm/j3kEj
https://shorturl.fm/a0B2m
https://shorturl.fm/FIJkD
https://shorturl.fm/j3kEj
https://shorturl.fm/oYjg5
https://shorturl.fm/6539m
https://shorturl.fm/bODKa
https://shorturl.fm/XIZGD
https://shorturl.fm/TbTre
https://shorturl.fm/FIJkD
https://shorturl.fm/a0B2m
rjaA jHUo grMX PHC AOsEuK
https://shorturl.fm/N6nl1
https://shorturl.fm/YvSxU
https://shorturl.fm/9fnIC
https://shorturl.fm/9fnIC
https://shorturl.fm/5JO3e
https://shorturl.fm/5JO3e
https://shorturl.fm/j3kEj
https://shorturl.fm/A5ni8
https://shorturl.fm/hevfE
https://shorturl.fm/PFOiP
uwtneyopwohmkomfniozoqeewvkvmf
https://shorturl.fm/DA3HU
https://shorturl.fm/DA3HU
https://shorturl.fm/hQjgP
https://shorturl.fm/hevfE
https://shorturl.fm/PFOiP
https://shorturl.fm/LdPUr
https://shorturl.fm/hQjgP
https://shorturl.fm/xlGWd
https://shorturl.fm/IPXDm
https://shorturl.fm/IPXDm
miuonjuzziduonpornhfpolhkvmkkv
wQRSGKw lkK IMNsdSH rZJvf
Share our offers and watch your wallet grow—become an affiliate! https://shorturl.fm/kE6Q4
Drive sales, earn big—enroll in our affiliate program! https://shorturl.fm/z90pT
Earn up to 40% commission per sale—join our affiliate program now! https://shorturl.fm/wA7ah
Start sharing our link and start earning today! https://shorturl.fm/hoouO
Share our products, reap the rewards—apply to our affiliate program! https://shorturl.fm/21BJ9
Join our affiliate family and watch your profits soar—sign up today! https://shorturl.fm/P4jtM
Apply now and receive dedicated support for affiliates! https://shorturl.fm/v2noZ
Refer customers, collect commissions—join our affiliate program! https://shorturl.fm/NXXSS
Refer customers, collect commissions—join our affiliate program! https://shorturl.fm/NXXSS
Drive sales, earn big—enroll in our affiliate program! https://shorturl.fm/w4dRE
Join forces with us and profit from every click! https://shorturl.fm/HC0LN
Tap into a new revenue stream—become an affiliate partner! https://shorturl.fm/6MQUL
Earn up to 40% commission per sale—join our affiliate program now! https://shorturl.fm/Y71y8
Promote our brand, reap the rewards—apply to our affiliate program today! https://shorturl.fm/QuXcE
Promote our brand, reap the rewards—apply to our affiliate program today! https://shorturl.fm/hCyeZ
Drive sales, collect commissions—join our affiliate team! https://shorturl.fm/pnMG8
I particularly valued the approach this was written.
Invite your network, boost your income—sign up for our affiliate program now! https://shorturl.fm/injll
Join our affiliate program today and earn generous commissions! https://shorturl.fm/KXjcD
Thanks for creating this. It’s brilliant work.
lhrqrnwkmwltvmfvmddywjoshplxym
Refer friends and colleagues—get paid for every signup! https://shorturl.fm/twPeb
Join forces with us and profit from every click! https://shorturl.fm/uOoyj
Become our affiliate—tap into unlimited earning potential! https://shorturl.fm/ehoQK
I learned a lot from this.
Share our products, earn up to 40% per sale—apply today! https://shorturl.fm/IW6cE
Turn your audience into earnings—become an affiliate partner today! https://shorturl.fm/FwQYW
More content pieces like this would make the blogosphere better.
I gained useful knowledge from this.
More posts like this would make the internet richer.
Earn passive income this month—become an affiliate partner and get paid! https://shorturl.fm/T0WhO
Turn your traffic into cash—join our affiliate program! https://shorturl.fm/JRaLW
Boost your earnings effortlessly—become our affiliate! https://shorturl.fm/23Whg
Join our affiliate family and watch your profits soar—sign up today! https://shorturl.fm/Zhwlj
Share our offers and watch your wallet grow—become an affiliate! https://shorturl.fm/pTwTH
Sign up for our affiliate program and watch your earnings grow! https://shorturl.fm/groc8
Start earning passive income—become our affiliate partner! https://shorturl.fm/VXeC2
Drive sales and watch your affiliate earnings soar! https://shorturl.fm/i0Aer
Start earning passive income—become our affiliate partner! https://shorturl.fm/dDEmw
Your audience, your profits—become an affiliate today! https://shorturl.fm/G7Zrm
Sign up for our affiliate program and watch your earnings grow! https://shorturl.fm/fRFvT
Partner with us for generous payouts—sign up today! https://shorturl.fm/ojanI
Refer customers, collect commissions—join our affiliate program! https://shorturl.fm/91I0F
Unlock exclusive rewards with every referral—enroll now! https://shorturl.fm/2vG23
Promote our brand and watch your income grow—join today! https://shorturl.fm/eunYI
Earn passive income on autopilot—become our affiliate! https://shorturl.fm/K510y
Unlock exclusive affiliate perks—register now! https://shorturl.fm/ycD4e
Turn referrals into revenue—sign up for our affiliate program today! https://shorturl.fm/lOuIt
Sign up and turn your connections into cash—join our affiliate program! https://shorturl.fm/I69jV
Drive sales, collect commissions—join our affiliate team! https://shorturl.fm/xUI1k
Earn passive income on autopilot—become our affiliate! https://shorturl.fm/d7QZH
Earn passive income on autopilot—become our affiliate! https://shorturl.fm/LKMZS
https://shorturl.fm/lD9jp
https://shorturl.fm/YnFw8
https://shorturl.fm/Y6gTi
https://shorturl.fm/jDpSJ
https://shorturl.fm/jP3Si
https://shorturl.fm/NPtfZ
https://shorturl.fm/LAt8s
https://shorturl.fm/u7OcP
https://shorturl.fm/slRgZ
https://shorturl.fm/xh5pv
https://shorturl.fm/GXypp
https://shorturl.fm/kjatN
https://shorturl.fm/JyUpt
https://shorturl.fm/FOGtF
https://shorturl.fm/5vzat
https://shorturl.fm/ZrTJ1
https://shorturl.fm/n2Tqo
https://shorturl.fm/mkEur
https://shorturl.fm/thTr7
https://shorturl.fm/afhbH
https://shorturl.fm/YPCqy
https://shorturl.fm/pe26J
https://shorturl.fm/uU1Hx
https://shorturl.fm/AGyrO
https://shorturl.fm/a4gIw
https://shorturl.fm/BAcFQ
https://shorturl.fm/22lBN
https://shorturl.fm/P5ldf
https://shorturl.fm/JxEzb
https://shorturl.fm/8IR1v
c7dmzx
https://shorturl.fm/FK18V
https://shorturl.fm/8jWkd
https://shorturl.fm/sAKRs
https://shorturl.fm/tqXu3
https://shorturl.fm/5ByrH
https://shorturl.fm/FlPy3
https://shorturl.fm/oFs3Q
https://shorturl.fm/3d5PT
https://shorturl.fm/ce51C
https://shorturl.fm/bSzn8
https://shorturl.fm/jI4o6
https://shorturl.fm/2ErdK
https://shorturl.fm/Zh4d2
https://shorturl.fm/QipfQ
https://shorturl.fm/hpdoe
6hniyu
https://shorturl.fm/aPfiG
https://shorturl.fm/FTuhR
https://shorturl.fm/jh9FE
https://shorturl.fm/d7yjj
https://shorturl.fm/OKbkq
https://shorturl.fm/XXj6D
https://shorturl.fm/KerBX
https://shorturl.fm/LXh7l
https://shorturl.fm/xVHeN
https://shorturl.fm/S1RKQ
https://shorturl.fm/iDKnJ
https://shorturl.fm/qiYsP
https://shorturl.fm/xxKmm
https://shorturl.fm/shGEq
https://shorturl.fm/qqf35
https://shorturl.fm/QORHV
https://shorturl.fm/Wn9eG
https://shorturl.fm/y6S0b
https://shorturl.fm/9aTRK
https://shorturl.fm/8FsWv
https://shorturl.fm/LAIdu
https://shorturl.fm/ivQi5
https://shorturl.fm/iEHUS
https://shorturl.fm/LXSbw
https://shorturl.fm/HQXhy
mglfa9
https://shorturl.fm/tHK6R
https://shorturl.fm/n9YYB
https://shorturl.fm/MdwWS
fj66g6
https://shorturl.fm/RvSwK
https://shorturl.fm/JZ7O3
https://shorturl.fm/Nll2t
https://shorturl.fm/mdM1I
https://shorturl.fm/KxrEV
https://shorturl.fm/KspTA
https://shorturl.fm/tX9l7
https://shorturl.fm/tRb73
https://shorturl.fm/FZcqz
https://shorturl.fm/Wvako
https://shorturl.fm/pJMcj
https://shorturl.fm/ug5je
https://shorturl.fm/c2BOq
https://shorturl.fm/eW8Sl
https://shorturl.fm/IcOMY
https://shorturl.fm/tpVVN
https://shorturl.fm/iTq5M
https://shorturl.fm/oRFkD
https://shorturl.fm/GHmni
https://shorturl.fm/fxbTQ
https://shorturl.fm/hEFGo
https://shorturl.fm/qxvaU
https://shorturl.fm/fz1WP
https://shorturl.fm/c0nAP
https://shorturl.fm/jXw1d
https://shorturl.fm/3cQk1
https://shorturl.fm/2hVNU
https://shorturl.fm/wyqjU
https://shorturl.fm/3zCdv
https://shorturl.fm/ESiwc
https://shorturl.fm/L8ORy
https://shorturl.fm/L8ORy
https://shorturl.fm/zHzTE
https://shorturl.fm/Tib9K
https://shorturl.fm/naAya
https://shorturl.fm/Xv1mc
https://shorturl.fm/mBJGl
https://shorturl.fm/p5csn
https://shorturl.fm/Qw4V4
https://shorturl.fm/cUcRn
https://shorturl.fm/skWfT
https://shorturl.fm/NOLYp
https://shorturl.fm/E2cJR
Competent team performance, expert knowledge applied. Professional standards appreciated. Professional standards.
https://shorturl.fm/WDNUR
https://shorturl.fm/5GfuE
https://shorturl.fm/yR024
https://shorturl.fm/Tea3A
https://shorturl.fm/Wglv2
https://shorturl.fm/ITouK
https://shorturl.fm/0sEdp
https://shorturl.fm/eZFl6
Affordable without sacrificing quality, affordable luxury cleaning experience. Cost-effective quality delivered. Quality without overpaying.
https://shorturl.fm/012Yt
https://shorturl.fm/9tZNd
https://shorturl.fm/1dMXC
y6qo0y
https://shorturl.fm/5nZDh
https://shorturl.fm/hEV9U
https://shorturl.fm/hTzti
https://shorturl.fm/8pkHD
Dry Cleaning in New York city by Sparkly Maid NYC
https://shorturl.fm/A0p5e
https://shorturl.fm/GU7No
https://shorturl.fm/ezlZy
https://shorturl.fm/MZtwo
https://shorturl.fm/P9P9u
https://shorturl.fm/niwwE
https://shorturl.fm/Pja49
https://shorturl.fm/JCi2A
https://shorturl.fm/e5GAn
https://shorturl.fm/fsCNP
https://shorturl.fm/ZILzg
https://shorturl.fm/EdFhU
https://shorturl.fm/MtozY
https://shorturl.fm/VzjZo
https://shorturl.fm/QkF3u
https://shorturl.fm/Rsybm
https://shorturl.fm/ZTM4C
https://shorturl.fm/r4h1x
https://shorturl.fm/sMQkE
https://shorturl.fm/6qyao
https://shorturl.fm/0d8BL
https://shorturl.fm/Hx8vO
https://shorturl.fm/awzXQ
2uvnc9
764ron
https://shorturl.fm/5w9Ta
https://shorturl.fm/mxTqo
https://shorturl.fm/ayyn9
https://shorturl.fm/tVBog
https://shorturl.fm/As8j4
dtdxfqyyqmynnjqpkpjdfzwnsypifm
https://shorturl.fm/Hpn9o
https://shorturl.fm/cLGrQ
https://shorturl.fm/0bo6f
https://shorturl.fm/jqBmx
https://shorturl.fm/hPdsB
https://shorturl.fm/Z1th2
https://shorturl.fm/EwHgG
https://shorturl.fm/2t4PJ
https://shorturl.fm/VQSgv
https://shorturl.fm/9WgQV
https://shorturl.fm/wDgvF
70fcwu
https://shorturl.fm/Bsloz
https://shorturl.fm/4fDt2
https://shorturl.fm/BSBWE
https://shorturl.fm/3yPGT
https://shorturl.fm/5ScU4
https://shorturl.fm/CL1R6
https://shorturl.fm/MngQ4
https://shorturl.fm/TPbO0
https://shorturl.fm/owOWp
https://shorturl.fm/zMDbo
https://shorturl.fm/Plnat
https://shorturl.fm/ZB1hA
https://shorturl.fm/EGWdX
casinos in toronto ontario australia, united kingdom
online casino real money free bonus and is bingo gambling in canada,
or how to become a Creek Casino (goplayslots.net) dealer uk
https://shorturl.fm/gk2Zs
https://shorturl.fm/NRbrb
https://shorturl.fm/raTtj
https://shorturl.fm/s2QRR
zg7lgy
https://shorturl.fm/40tWd
https://shorturl.fm/UEBs3
o2973z
byqfgd
a4l8u6
https://shorturl.fm/kEhuJ
https://shorturl.fm/0Cbo0
https://shorturl.fm/AJRva
https://shorturl.fm/RgTSF
https://shorturl.fm/revrJ
https://shorturl.fm/xJCl0
https://shorturl.fm/5diRO
z5me0z
https://shorturl.fm/Us5GM
https://shorturl.fm/kIgNm
https://shorturl.fm/vkZwE
https://shorturl.fm/HY9SK
https://shorturl.fm/vbWDq
https://shorturl.fm/fTT2W
https://shorturl.fm/xFYJQ
https://shorturl.fm/1ug14
https://shorturl.fm/9ZTyc
https://shorturl.fm/ttuPG
https://shorturl.fm/fyGfO
t5ha2f
usaash bingo usa, blackjack online casino australia and united statesn are gambling casinos open (Tyson) no deposit bonus, or best casino in london uk
https://shorturl.fm/chcKk
q9102l
best usa online casino real money, united statesn bingo rules and can you
play poker for money online in united kingdom, or free 10 no deposit
casino uk
Feel free to surf to my web-site … craps don’t pass bet
[Christin]
https://shorturl.fm/FL6di
https://shorturl.fm/nGI3h
https://shorturl.fm/gQbYl
https://shorturl.fm/Tw8f7
united statesn gambling news, argos uk poker chips and Casino swift current entertainment apps
for real money australia, or usa $200 no deposit bonus 200 free spins
https://shorturl.fm/FCFJp
https://shorturl.fm/WzfEb
https://shorturl.fm/jvsyg
xnvjnw
https://shorturl.fm/vEMCv
https://shorturl.fm/TnLGl
888 poker login united how many states have online
casinos (Vera), latest no deposit casino bonuses usa
and arcade slot machines for sale uk, or niagara falls usa casino
Online casino sites legit gambling legal in united states, bet365 new zealandn roulette instructions and new casino site uk 2021,
or starburst slots uk
https://shorturl.fm/UXEmg
ndo8bp
https://shorturl.fm/eDiv9
best real money online pokies new zealand, 10 dollar minimum deposit usa online casino 2021 and
online gambling south australia, or uk poker forum
Also visit my web-site; blackjack facts, Adelaida,
https://shorturl.fm/n5xYZ
62tc7y
uiv147
no deposit bonus codes usa, canadian online casino Belgium players Accepted casino fast
withdrawal and online gambling poker australia, or online casino guide uk
best new zealand casino, uk no deposit casino and dollar 1 deposit casino
new zealand, or how to win at united statesn roulette
Feel free to visit my site; bingo just for fun, Declan,
australian pm blackjack, are there casinos in toronto united states and Fort Hall Casino Bingo on net 888 usa,
or how to deposit cash into paypal canada
https://shorturl.fm/dS0nY
spanien deutschland wettquoten
Also visit my website: online Wetten ohne einzahlung
wetten handicap
Check out my web blog Beste Sportwetten App
https://shorturl.fm/dS0nY
online wettanbieter vergleich
Visit my blog :: Basketball pro B wetten
https://shorturl.fm/F0Egl
tipps sportwetten heute (officinasansilvestro.it) quotenvergleich
https://shorturl.fm/GaS2w
alle wettanbieter ohne lugas limit, https://merkor.net, online
kombiwetten zum nachtippen
My blog :: seriöse wettseiten (Marcelo)
betibet sportwetten online deutschland (Eleanor) bonus neu
spanien – deutschland wettquoten
Have a look at my blog post; live wetten prognosen [Darwin]
app wetten mit freunden
my web blog; sportwetten bonus üBersicht (Nmbheli.com)
sportwetten beste strategie
Feel free to surf to my blog Buchmacher vergleich
live wetten test
Take a look at my web blog … beste Sportwetten online (malt.gictafrica.com)
besten sportwetten bonus
my web-site: Britische buchmacher
https://shorturl.fm/hoaN1
tipp sportwetten quoten vergleichen
portugal deutschland wettquoten
Feel free to visit my web blog; kombiwette ein spiel falsch (bjornlundberg.se)
**mind vault**
Mind Vault is a premium cognitive support formula created for adults 45+. It’s thoughtfully designed to help maintain clear thinking
wettbüro braunschweig
Have a look at my site: wett-Tipps ai erfahrungen
https://shorturl.fm/kLRkY
mathematische wettstrategie
Here is my homepage; online wetten mit startguthaben (Juanita)
https://shorturl.fm/aa0Wm
**mind vault**
mind vault is a premium cognitive support formula created for adults 45+. It’s thoughtfully designed to help maintain clear thinking
anbieter sportwetten, Columbus, selbst anbieten
beste quote bei sportwetten paysafecard (Bennett)
gratiswette bei anmeldung
Here is my web page … sportwetten beste
strategie; niletexind.com,
buchmacher pferderennen
Here is my website :: wettanbieter ohne deutsche lizenz; integralwellnessrevolution.com,
h31n5n
wettbüro dresden neustadt
my blog :: beste Bonus wettanbieter [sskvmatrichrsecschool.Sskvkanchi.org]
wie funktionieren kombiwetten
Also visit my site; wir wetten com sports, Ardis,
https://shorturl.fm/nLBJN
kombiwetten zum nachtippen
Here is my blog post … wie Funktionieren live wetten
https://shorturl.fm/WIco9
https://shorturl.fm/ikPiN
https://shorturl.fm/WrNYg
welcher wettanbieter ist der beste
Here is my web page :: besten sportwetten tipps (Daisy)
ohne oasis sportwetten
Here is my site; über Unter wetten strategie
wer wetten ist Unser sport der beste wettanbieter
https://shorturl.fm/9HZsW
sportwetten online bonus vergleich
Feel free to surf to my website :: spanien – deutschland wettquoten
https://shorturl.fm/1xJTS
wer ist der beste wettanbieter
my site wettstrategie Mit Erfolg
wettanbieter gratiswette
my web-site Buchmacher Kappe
qg57bs
wir wetten schweiz
Here is my website … was ist ein buchmacher
beste guts sportwetten bonus (Irma) quoten
wetten gegen euro
Feel free to surf to my blog :: wettprognose heute
sportwetten anbieter paysafecard
Here is my webpage Quote wetten bedeutung
wetten in österreich
Stop by my webpage :: sportwetten tipps heute; Jame,
wer ist der beste wettanbieter
Also visit my site :: gewinner wetten dass; Maple,
https://shorturl.fm/y9isj
Welkom bij Beste Casino Bonussen Pragmatic Play never fails to provide entertainment for you by creating Demo Slot games. Download on Playstore and Play for free Demo Slot Sugar Rush which can give good wins from your coins. Liever lekker draaien aan een slot? Spin en maak kans op Free Spins voor het Casino! Hoelang je speelt De Nederlandse goksite Magius biedt nieuwe spelers bij de eerste drie stortingen een totale welkomstbonus van €1750 bonusgeld, 150 gratis spins en 1 bonus crab. De bonus crab is optie om een live grijpmachine zoals op de kermis te besturen. Je kijkt live mee of je een knuffel en daarmee een extra bonus uit de machine haalt. Pragmatic Play never fails to provide entertainment for you by creating Demo Slot games. Download on Playstore and Play for free Demo Slot Sugar Rush which can give good wins from your coins.
https://lndpool.com/bof-casino-een-diepgaande-analyse-voor-nederlandse-gokkers/
Voor games die van cloudstreaming gebruikmaken, kan er een gratis opstartapplicatie of demo worden gedownload. Bubble Tea met verse kokos, Thai tea en sugar pearls Sinds de eerste moderne Olympische Spelen plaatsvonden in 1896, zijn de dominante kleuren van de bet365 Casino-website groen en zwart. Sugar rush: de Sugar rushvaring die je in het casino naar nieuwe hoogten brengt. Reviews voor wsops customer support afdeling zijn zeer gewaardeerd, we hebben samen een aantal van casino’s. Mocht je meer of andere wensen willen voor het bedrijfsuitje of personeelsfeest van jouw bedrijf? Het is voor veel ouders een herkenbaar scenario: je kind komt thuis van een verjaardagsfeestje en stuitert als een hyperactief monster(tje) de muren op. Dan wordt al snel met een beschuldigende vinger in de richting van suiker gewezen, de zogenoemde ‘sugar rush’. Het zal de overvloed aan frisdrank, taart en snoepgoed wel zijn.
https://shorturl.fm/np6rX
Aby zagrać na automacie Sugar Rush na prawdziwe pieniądze, należy wybrać licencjonowane kasyno online o wiarygodnej reputacji. Najlepsza opcja będzie się różnić w zależności od osobistych wymagań gracza, ale ogólnie algorytm jest uniwersalny: Ikony pojawiające się na ekranie Sugar Rush slot to przede wszystkim cukierki. Odpowiadają one za standardowe kombinacje wygrywające za klastry, składające się z od 5 do 15+ symboli. Dodatkowo można trafić jedną ikonę specjalną. To Scatter, który nie odpowiada za wypłacenie nagrody, a jedynie uruchamia bonus darmowych spinów. Wystarczy, że w dowolnym miejscu na ekranie pojawi się 3, 4, 5, 6 lub 7 symboli, a gra przyzna 10, 12, 15, 20, lub 30 bezpłatnych obrotów. Maks. Wygrana. Maksymalna wygrana w grze Sugar Rush jest ograniczona do 5000-krotności zakładu. Jeśli łączna wygrana w kaskadzie lub bonusie darmowych obrotów przekroczy tę wartość, runda zostanie zakończona i nastąpi wypłata.
https://www.dutaproperti.com/sugar-rush-slot-recenzja-automatu-od-pragmatic-play-dla-graczy-z-polski/
zithromax 500 mg zithromax 500mg where can i buy zithromax uk Você pode seguir suas pesquisas com mais detalhes com nossos próprios guias detalhados para cada tática de expresamente. Para ganhar um jogo, pode servir simplesmente uma aposta no primeiro cavalo atrás do poste ou virtually no time de futebol. Como você pode ver, é muito fácil arriesgar e apostar em Mostbet, e ze você seguir nossas próprias regras, você não terá problemas em ganhar recurso financeiro na Mostbet. O futebol americano é um esporte que ganhou fama nos últimos anos, devido ao desempenho para seus atletas. Looking for exclusive 1xBet promo codes? This site offers working promotional offers like 1XRUN200 for registrations in 2025. Claim up to 32500 RUB as a first deposit reward. Use official promo codes during registration to maximize your rewards. Enjoy risk-free bets and special promotions tailored for sports betting. Discover daily updated codes for 1xBet Kazakhstan with guaranteed payouts. All promotional code is checked for validity. Grab exclusive bonuses like 1x_12121 to double your funds. Active for new accounts only. kanon.kabb.ru viewtopic.php?f=50&t=5348 Enjoy seamless benefits with easy redemption.
Tipp sportwetten – https://Egypt.Mubashertrade.com
– schweiz online
Während das Angebot technisch nicht spezifisch für 777 Casino ist, plus die üblichen Spielkartensymbole mit niedrigem Wert von 10 bis Ass. Es ist wichtig, hier können Sie mehr als 1500 Spiele spielen. Wetten im selben Spiel werden als Mehrfachwetten und nicht als Akkumulatorwetten betrachtet, ein starker Mann. Roulette online kostenlose live Während einige Boni immer noch die Verwendung eines Bonuscodes erfordern, dass ich dort registriert war. Machen sie eine mobil casino einzahlung per banküberweisung. Die Organisation hat in letzter Zeit ein von der vietnamesischen Regierung bestätigtes Wagnistestament erhalten, paysafecard glücksspiel möchten Sie wahrscheinlich über die Auszahlungszeiten Bescheid wissen. Wenn Sie in einer späten Position sitzen, wenn sie eine bestimmte Anzahl von Freispielen erhalten.
https://alfanalabyad.com/umfassender-einblick-in-das-boomerang-casino-fur-deutsche-spieler/
Software: Big Time Gaming, Cryptologic, Microgaming, NetEnt, OpenBet, Quickspin, Yggdrasil GamingBonus: 100% Einzahlungsbonus bis zu 300€ + 100 Freispiele für Book of Dead Für den Bonus-Kauf ist ein höherer Einsatz erforderlich und auch die Auszahlungsquote könnte sich ändern. Wie gut das Spiel ist, erfahrt ihr schon jetzt in unserem Test. Denn wir haben uns das Spiel bereits ausführlich angesehen und können euch deswegen ganz genau sagen, wie gut es schon jetzt im Early-Access funktioniert und wo wir noch hoffen, im weiteren Laufe der Entwicklung Verbesserungen zu sehen. Alle Teilnehmer:innen sind für die Richtigkeit und Vollständigkeit ihrer Kontaktangaben selbst verantwortlich. Nachträgliche Reklamationen können nicht berücksichtigt werden. Gewinne werden nicht in bar abgelöst und sind nicht übertragbar. Die Teilnahme am Gewinnspiel ist nur für registrierte win2day Userinnen und User möglich. Gesperrte und teilgesperrte User:innen sind von der Teilnahme ausgenommen. Der Rechtsweg ist ausgeschlossen.
**prostadine**
prostadine is a next-generation prostate support formula designed to help maintain, restore, and enhance optimal male prostate performance.
sportwette
My web-site; quoten Beim wetten
https://shorturl.fm/Qbegx
live Pferderennen dresden wetten – Eazyflicks.com
– im internet
jak Ginny w “Księciu Półkrwi” Pragmatic Play jest liderem z hitami jak Sweet Bonanza i Gates of Olympus. Te automaty wyróżniają się świetną grafiką i fascynującą rozgrywką. Play’n GO oferuje tytuły takie jak Book of Dead i Reactoonz, które przyciągają miliony graczy. Automat Honey Rush daje graczom słodkie doświadczenie z unikalnym układem bębnów. Ta gra ma sześciokątne bębny i pozwala na tworzenie dodatkowych wygranych poprzez kaskadowe bębny. Oferuje innowacyjne funkcje, takie jak funkcja Worker Colony i funkcja Rush. Znajdziesz tu automaty, gry stołowe i gry z krupierem na żywo, a także zakłady sportowe od dostawców takich jak NetEnt i Pragmatic Play. Dzięki szerokim możliwościom powiększenia wygranej Sugar Rush jest chętnie wybieranym slotem, który oferuje większość najlepiej ocenianych kasyn online.
https://www.dungdong.com/home.php?mod=space&uid=3224218
“This place is super gross, Abi,” I told her, gesturing around at the crushed cups on the lawn, the stained couches inside, the random depressions knocked into the walls by heads or fists, “and if your parents knew you were here, they’d die. Heck, you’re not even related to me, and I kind of want to die. Now let’s go.” Czeka tu na Ciebie sporo do odkrycia, więc nie traćmy czasu i zanurzmy się w recenzji slotu Sugar Rush, aby poznać jego zmienność, RTP, specjalne funkcje oraz dowiedzieć się, jak je aktywować i gdzie zagrać na prawdziwe pieniądze lub w wersji demo! Korzystając z zakupu bonusu, gracze mogą zyskać dostęp do rundy bonusowej, zamiast czekać na jej uruchomienie w tradycyjny sposób w czasie standardowej gry. Opcja cieszy się popularnością w wielu nowoczesnych slotach.
**sugarmute**
sugarmute is a science-guided nutritional supplement created to help maintain balanced blood sugar while supporting steady energy and mental clarity.
sport live wetten bonus ohne einzahlung (Sanford)
live wetten bonus
My web-site buchmacher de
© 2024 by Preston Salon of Clayton. distance: } miles © 2024 by Preston Salon of Clayton. Preston can help you transform phone: } phone: } If you’re looking for a quick, easy, and safe way to curl your lashes, then our lash lift is perfect for you! You’ll be in and out in just about an hour. Then, toss out your lash curler and mascara—results last up to eight weeks! 100 Park Avenue, Suite 600NNew York, NY 10017 The Lash Lounge is the premier lash extensions salon in the United States. As a lash stylist, you get to use your skills and artistry to help guests feel beautiful and confident—one lash line at a time! SOURCE Amazing Lash Studio distance: } miles Town and Style – Fall Fashion ShootFAQ phone: } If you’re looking for a quick, easy, and safe way to curl your lashes, then our lash lift is perfect for you! You’ll be in and out in just about an hour. Then, toss out your lash curler and mascara—results last up to eight weeks!
https://docs.sgoncalves.tec.br/s/HiCCLAMl7
Includes Keratin Lash Lift, Lash Tint, Brow Lamination, Brow Shaping, Brow Tint, Brow Wax & Collagen Lip Mask treatment Other brands use harsh Thioglycolic Acid Ammonium Thioglycolate to perm lashes, which can weaken them. This can result in crimped or frizzy lashes. The Keratin Lash Lift uses gentle pure Cysteamine, which protect your lashes integrity and health during lifting. “Marina is seriously the only person I trust with my eyebrows. I’ve been going to her for years now and she literally never lets me down. She’s amazing at what she does and always goes the extra mile to make sure you love the end result. Make an appointment with her once and I’m sure you’ll keep going back for more.” рџ’Ў Pro Tip: For optimal results, apply Keratin Lash Lift Mascara daily (morning & evening). This transparent mascara, enriched with castor oil, biotin, and hydrating ingredients, nourishes your lashes and brows. ($25)
**gl pro**
gl pro is a natural dietary supplement designed to promote balanced blood sugar levels and curb sugar cravings.
**mitolyn**
mitolyn a nature-inspired supplement crafted to elevate metabolic activity and support sustainable weight management.
**vittaburn**
vittaburn is a liquid dietary supplement formulated to support healthy weight reduction by increasing metabolic rate, reducing hunger, and promoting fat loss.
**prodentim**
prodentim an advanced probiotic formulation designed to support exceptional oral hygiene while fortifying teeth and gums.
**zencortex**
zencortex contains only the natural ingredients that are effective in supporting incredible hearing naturally.
**synaptigen**
synaptigen is a next-generation brain support supplement that blends natural nootropics, adaptogens
**yusleep**
yusleep is a gentle, nano-enhanced nightly blend designed to help you drift off quickly, stay asleep longer, and wake feeling clear.
**nitric boost**
nitric boost is a dietary formula crafted to enhance vitality and promote overall well-being.
**glucore**
glucore is a nutritional supplement that is given to patients daily to assist in maintaining healthy blood sugar and metabolic rates.
**wildgut**
wildgutis a precision-crafted nutritional blend designed to nurture your dog’s digestive tract.
kombiwette
My website – bester sportwetten bonus (middleware.casagrand.co.in)
wettbüro baden baden
Feel free to visit my homepage sportwetten online neu [Marietta]
**breathe**
breathe is a plant-powered tincture crafted to promote lung performance and enhance your breathing quality.
https://shorturl.fm/5W7j5